The discovery of restriction endonucleases
|
|
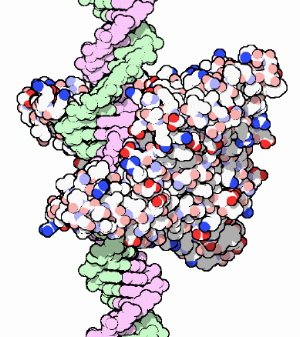 Restriction endonucleases bind and cut DNA. Here, the restriction endonuclease EcoRI is bound to a (lavender and green) DNA helix.
Restriction endonucleases bind and cut DNA. Here, the restriction endonuclease EcoRI is bound to a (lavender and green) DNA helix.
For their 1970 discovery of restriction endonucleases (often called by the shorter name restriction enzymes) Werner Arber, Hamilton Smith, and Daniel Nathans received the 1978 Nobel Prize for Physiology or Medicine. HindII was the first restriction enzyme to be isolated, but many others were later discovered and characterized.
Restriction enzymes are produced by bacteria, in which they function as a simple immune system — A restriction enzyme protects the bacterium producing it from viral infection by chopping up invading viruses, which are composed of either RNA or DNA.
Each restriction enzyme recognizes and cuts a specific, short nucleotide base sequence, not found in the genome of the bacterium that produces it. For example, the restriction enzyme EcoRI will bind a DNA helix (right) and cut it at (and only at) points where the nucleotide sequence GAATTC occurs (see lower figure at right). This specific 6-nucleotide sequence is the restriction site of EcoRI (other restriction enzymes have other restriction sites). The green line in the figure indicates the path of the cut created by the enzyme.
But restriction enzymes will cut not only the DNA and RNA of viruses invading a bacterium, but also DNA and RNA of any kind. This makes them incredibly useful for constructing composite DNA sequences and for carrying out molecular genetic investigations.
For example, restriction enzymes allowed the creation of a cheap and efficient process for the production of insulin. This was done by adding an insulin gene to a plasmid in the bacterium Escherichia coli. The altered bacterium was then used to mass produce the drug.