On the Origins of New Forms of Life
5.1: On the Prevalence of Polyploidy
(Continued from the previous page)
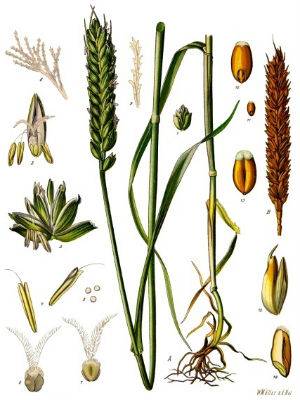 Wheat (Triticum aestivum), an organism resulting from polyploidization.
Wheat (Triticum aestivum), an organism resulting from polyploidization.
In connection with stabilization theory, it's important to realize that polyploid organisms are extremely common. An initial point to consider in assessing the prevalence of polyploidy is that biologists tend to assume any organism of unknown status is diploid. For example, Wu et al. (2001) comment that
Here "traditionally considered" means "considered in the absence of evidence to the contrary." This particular assumption therefore has the odd effect of leading one to suppose that most organisms are diploid (since the ploidy level of most organisms has not been investigated). In fact, however, it is unknown whether the typical organism is diploid.
Despite this traditional default assumption, it is now well established that the production of new forms of life through polyploidy is extremely common in plants. Masterson (1994) estimated that the rate of polyploidy among plants is between 30 and 80 percent. More than half a century ago, Stebbins (1950: 299) said it was already known that important crop plants such as wheat, oats, cotton, tobacco, potato, banana, coffee, and sugar cane were polyploids and that the actual parentage of wheat (Triticum aestivum), cotton (Gossypium hirsutum), and tobacco (Nicotiana tabacum) had already been determined. More recently, Hilu (1993) estimated that about 75 percent of domesticated plants are polyploid.
One of the most surprising findings of recent genetic research is the ubiquity of polyploidy — even in taxonomic categories where no one expected to find it. Over the last decade certain organisms have been singled out for intense genetic analysis with modern automated techniques. Detailed examination of an array of these so-called "model organisms" has shown that most are in fact polyploid. For example, rice (Oryza sativa) was recently demonstrated to be an ancient polyploid. The same is probably true for yeast (Saccharomyces cerevisiae).
Indeed, Soltis (2005: 5) reports it now appears all angiosperms are either polyploid or descended from polyploids. This means the vast majority of extant plants are polyploid — some 240,000 angiosperms are treated as species (among gymnosperms, the second largest plant category, only about 720 are so treated). Even more surprising is the finding that all vertebrates are probably also ancient polyploids. In other words, a doubling of chromosome number appears to have occurred during the evolution of the vertebrates at an early stage prior to their diversification. These findings represent quite a change in perspective. Writing only a generation ago, Schultz (1980: 314) remarked that a majority of biologists were still unaware that polyploid vertebrates even existed.
Ancient polyploids are often difficult to recognize because chromosomes fuse or break over time, hampering analyses based on chromosome counts. Also, mutations subsequent to the original event initially producing a polyploid may obscure an organism's true status. As a result, with the passing generations the chromosomes of a polyploid start to look more like those of a diploid as such changes accumulate, making chromosomes that were originally duplicates become more distinct from each other. Hence, even many types of organisms that now appear to be diploid must have been polyploid in origin.
The idea that polyploidy is rare among animals was once widely accepted. The arguments most often offered to support this claim were
-
that the extensive gene duplication seen in polyploids would dilute the effects of new mutations and so make significant adaptive changes unlikely;
and - that animals are somehow too "complex" to tolerate the dramatic genetic changes associated with polyploidy.
Neither idea is accepted today. The former has been dismissed because gene duplication is now seen as actually opening an opportunity for genic evolution since one gene copy can go on performing its accustomed function while other copies are freed to mutate and adapt to new functions. The latter objection, as Orr (1990) points out, has been invalidated by the fact that many animals are now actually known to be polyploid. Gregory and Mable (2005: 478), who recently surveyed known animal polyploids, note that polyploidy is
Among vertebrates, polyploidy is especially extensive among fishes, but there are numerous known examples, too, among amphibians and reptiles. In particular, it now appears that a doubling in chromosome number preceded the diversification of teleost fishes (superorder Teleostei), which means it is likely this entire group is polyploid. The vast majority of all fishes are teleosts. In fact, Teleostei is by far the most diverse vertebrate group, containing roughly half of all vertebrate taxa treated as species. The vast majority of all extant fishes are teleosts. We can therefore no longer speak of "the scarcity of polyploidy in the animal kingdom as a whole," as Stebbins (1950: 368) once did. Certain large families within Teleostei (e.g., Salmonidae, Catostomidae) are now known without doubt to be polyploid. Others (e.g., Cyprinidae) are known to contain many polyploid forms of recent origin. Even among the non-teleosts (a distinct minority of fishes fall into this category), many polyploids are known. Gregory and Mable (2005: 477) say an ancient doubling of chromosome number in
Elsewhere Mable (2004: 453) notes,
Here Mable refers to the assertions of H. J. Muller (1925), whose views concerning the rarity of polyploidy in animals were long influential (Appendix E spells out the shortcomings of the arguments offered by Muller to support his mistaken idea that animal polyploidy must, for genetic reasons, be rare). Also, says Mable (ibid: 453), proponents of the idea that animals are less often polyploid than are plants have focused
These first reports to which Mable refers were made by Gallardo et al. (1999, 2004), who discovered that the Plains Viscacha-Rat, Tympanoctomys barrerae, is tetraploid. They say (2004: 443) evidence "strongly suggests a hybrid derivation" for this rat. Gallardo et al. (ibid) also say the recently described Golden Viscacha-Rat (Pipanacoctomys aureus) appears to be a tetraploid of hybrid origin.
Thus, polyploidy is nearly ubiquitous in plants. Among animals, it appears the typical vertebrate has a set of chromosomes that is the product of one or more ancient polyploidization events partially masked by subsequent point and chromosomal mutations. Moreover, many types of animals treated as species are known to be derived from more recent polyploidization events, as are many invertebrate forms. It would be tedious to list here all known cases of animal polyploids, but the curious reader is referred to Gregory and Mable's (2005) excellent review.
Polyploidy and Hybridization. Because of the strong association between polyploidy and hybridization (see Section 4), the wide prevalence of polyploidy indicates natural hybridization, too, is widely prevalent. Allopolyploidy (which, recall, is polyploidy resulting from hybridization) has long been thought to be important in plant evolution because of the large number of known natural plant amphiploids treated as species, and, more recently, because it is thought to open opportunities for new gene regulation mechanisms. Many natural allopolyploids have been recreated artificially by crossing their parents.
Hybridization produces polyploids not only in plant crosses, as has long been realized, but also in crosses between animals. For example, Kobayasi and Hashida (1977) report triploid males in the F1 progeny produced by crossing two diploids, Goldfish (Carassius auratus) and Cruscian Carp (C. carassius). polyploidy in the Bulinus truncatus/tropicalis complex of African snails is the result of hybridization, and in the European freshwater limpet, Ancylus fluviatilis. Some evidence suggests that polyploids are produced more frequently when the hybridizing parents are more disparate. In his research on toad hybrids in family Bufonidae, Bogart found that
Many supposed autopolyploids are actually derived from hybridization, but are not recognized as such due to the official taxonomic treatment of the hybridizing forms. That is, they are products of hybridization between forms that are distinct, but treated as conspecific. Suomalainen et al. (1987: 100) state that polyploid vertebrates of known origin have generally "proven either to be species hybrids or hybrids between different cytological races [i.e., different chromosets] of a single species." Among plants many cases reported as examples of natural autopolyploidy involve hybrids between populations treated as distinct races. There can be large numbers of structural differences between related chromosets. There can also be only a few, or even one. All degrees of difference exist. This is one reason why it is generally hard to distinguish allopolyploids derived from hybridization between chromosets with very similar karyotypes from autopolyploids derived from multiplication of a single chromosome set. Therefore, many polyploids that look like they are the result of doubling or multiplying one and the same chromosome set (autopolyploids) may actually be allopolyploids. As Grant (1981: 306) points out, even among the few accepted case of natural autopolyploidy
Although breeders using artificial techniques do often produce autopolyploids, they seem even more often to produce allopolyploids. It has long been known that natural polyploids among plants are most often produced by the latter of these two methods, that is, as hybrids. Nearly fifty years ago Stebbins (1959: 237–238) stated that "the production of stable, true-breeding new species through doubling the chromosome number of a sterile interspecific hybrid, is now generally recognized as one of the commonest ways in which plant species arise." For example, Brochmann et al. (2005) found that, among 47 plant polyploids from Svalbard Island, all were of hybrid origin. Today it is realized most natural polyploids, not just among plants but polyploids of any kind, are derived from hybridization (i.e., they are allopolyploids). Ancient polyploids are thought, like recent polyploids, to have been mostly produced by allopolyploidy rather than autopolyploidy.
Not only is there good reason, then, to suppose hybridization is widespread among plants, but there is also strong evidence the typical vertebrate is derived anciently and/or recently from hybridization. This conclusion seems inevitable given facts already mentioned:
- polyploidy is usually triggered by hybridization;
- vertebrates in general are now thought to be ancient polyploids;
- many extant vertebrate taxa are known to have had their origins recently as polyploids (produced by hybridization);
and - even many non-polyploid vertebrates treated as species are known to have had a hybrid origin.
This conclusion is in direct opposition to the traditional view that the great majority of vertebrates are diploid and rarely of hybrid origin.
Agamospermy. Agamospermy is widespread in plants. Among animals, it is limited to parthenogenesis (agamospermy in which embryos develop from unfertilized eggs). Animal groups in which parthenogens occur include ribbon worms, rotifers, tardigrades, reptiles, amphibians, sea anemones, flatworms, polychaete worms, starfishes, and insects. Suomalainen et al. (1987: 175) say parthenogenesis is known from all major insect groups except dragonflies. White (1973: 700) notes that only about one in a thousand of the various animal forms treated as species are actually known to be capable of parthenogenetic reproduction. However, it is not possible to infer the true number capable of parthenogenesis on the basis of the number currently known. After all, every year more different kinds of animals are found to have this capability. For example, the Yellow-spotted Goanna, the varanid lizard (mentioned earlier), has long been treated as a species but was only recently (2005) found to be capable of parthenogenetic reproduction. The tendency to assume organisms are sexual and not agamospermous until they are proved to be agamospermous is widespread (this is another of the unsubstantiated default assumptions that tend to shore up neo-Darwinian theory). But in the present context, it is less important to determine the number of existing agamosperms than to note that
- agamosperms are organisms whose origins are easily determined and
- their origins, when known, are consistently through hybridization.
Section 4 gave examples of agamosperms known to be derived from stabilization processes triggered by hybridization. Since natural polyploids most often arise through hybridization (i.e., most are allopolyploids), and since agamospermy is strongly associated with polyploidy (Christopher et al. 1991: 333; Gregory and Mable 2005: 442), one would expect agamospermy, too, to be closely associated with hybridization and very often to result from it. Such is the case. One of the most obvious facts indicating agamosperms are often of hybrid origin is the finding that many are of low sexual fertility and produce many bad gametes (a characteristic typical of hybrids). For example, many plant agamosperms are sexually sterile, or have reduced fertility, due to disrupted meiosis. There is no reason to expect low fertility to be a common characteristic of organisms capable of agamospermous reproduction if they are not commonly of hybrid origin.
Another clear indication of their hybrid origin is the observation that when agamosperms do reproduce sexually, they usually produce highly variable offspring, but not when they reproduce agamospermously. This hypervariability seen in the sexual reproductive mode is exactly like that seen in later-generation hybrids and therefore suggests that the parental plants are themselves hybrids. As Grant (1981: 424) notes, "Wide segregation of this sort as observed in [the agamosperms of] Potentilla, Rubus, Sorbus, and Citrus reveals the hybrid constitution of the agamospermous mother plant. The breeding behavior of the agamospermous plant, in short, is like that of an interspecific hybrid." Variation of this type in the F1 generation of hybrids produced by crossing the lemon (Citrus limon) and grapefruit (C. paradisi), which are both agamosperms, suggests that one or both of these common citruses are themselves of hybrid origin (recall that an F1 generation derived from hybridization between pure types normally is not variable).
The fact that agamosperms are very frequently the products of hybridization has long been recognized. Those of known origin are typically allopolyploids. Thus, Asker and Jerling (1992: 112–113) point out that many agamosperms "are of hybrid origin and have arisen as allopolyploids from sexual parents." For example, agamospermy has been shown to arise in hybrids involving the genera Hordeum and Triticum. For example, Mujeeb-Kazi (1981) produced hybrids between Hordeum vulgare (barley) and Triticum turgidum (rivet wheat) and between H. vulgare and T. aestivum (common wheat). He then backcrossed each of these two types of F1 hybrids to their respective wheat parents. Both backcrosses produced agamospermous plants, although none of the parents are agamosperms. Agamosperms are also known to have been produced by hybridization in the genera Antennaria, Calamagrostis, Crepis, Parthenium, Potentilla, and Rubus. Naturally occurring agamosperms in some of these genera (Antennaria, Potentilla, and Rubus) have been resynthesized by crossing their parents. Nogler (1984) says a cross between two sexual grasses, Schedonorus pratensis x S. phoenix (meadow fescue x tall fescue), produces agamospermous hybrids. Nearly all the agamospermous ferns studied by Manton (1950, ch. 11) were shown to be of hybrid origin. We have already encountered a specific example of agamospermy arising through hybridization: The fertile hybrid between cabbage and radish, radicole (Raphanobrassica), whose mode of origin has already been described, can reproduce sexually, vegetatively, and agamospermously. It is not only sexually fertile like its parents, but also produces seed without fertilization. So the cross producing this hybrid gave rise to a new stable form capable of agamospermous reproduction, whereas its parents are capable only of sexual and vegetative reproduction. The Maiden Campeloma (Campeloma parthenum), which, recall, is an allopolyploid parthenogenetic snail, is a population composed of multiple clones, each separately produced by separate hybridizations between its sexual parents.
Christopher et al. (1991: 344–345) state that "in the Maloideae [a subfamily of the rose family Rosaceae] interspecific hybridization provides a pathway for the generation of new taxonomically recognizable entities. Even in Pyrus, which is mostly diploid and sexual, hybridization is 'undoubtedly involved in the evolution of the genus' (Bell and Hough 1986). Polyploidy, apomixis, and self-compatibility may arise after hybridization and help isolate hybrids from parental species." They go on to give the genus Sorbus as an example (Christopher et al. use apomixis in the narrow sense of agamospermy, as do many other authors):
For example, they say, Sorbus aucuparia (European mountain ash) presumably crossed with S. rupicola (rock whitebeam), a tetraploid, to yield triploid S. arranensis (Arran whitebeam), which then crossed with S. torminalis (checkertree) to produce the tetraploid S. intermedia (Swedish Whitebeam). They also say, S. aucuparia and S. rupicola, the same pair that yielded S. arranensis on the Isle of Arran in Scotland, crossed to produce two other forms treated as species (S. minima in Wales and S. tamamsjanae in Armenia). These three hybrid forms, derived from the same cross, differ in morphology.
Vertebrate parthenogens of known origin are apparently all of hybrid origin. By chromosomal, molecular genetic, and morphological criteria, seemingly all types of parthenogenetic lizards thus far investigated have been shown to be of hybrid origin. For example, five lizards of the genus Lacerta are known to be of hybrid origin: L. armeniaca, L. dahli, L. rostombekovi, L. unisexualis, and L. uzzelli. They are all parthenogens derived from various crosses between sexual congeners. In point of fact, no matter the category of organism, virtually all agamosperms of known origin, whether animal, plant, or fungal, are derived from hybridization. The only exceptions that the writer has been able to identify are new forms produced by artificial techniques such as application of chemicals or heat shock. Some cases of this sort are discussed in Appendix C.
The origins of so many agamosperms are known because hybridity is easier to detect in these organisms than in the case of nonpolyploid sexual organisms derived from hybridization (i.e., recombinant derivatives). This is because in agamosperms meiotic recombination generally does not occur. The karyotype is therefore usually a stable combination of unaltered chromosomes anciently derived from the initial cross. This lack of alteration makes it easy to identify the parents involved in the original cross. Agamosperms, then, constitute a class of organisms that are
- commonly treated as species;
- often of known origin; and
- are typically derived from hybridization.
Clearly, it is not usual for agamosperms to come into being through gradual divergence under the influence of natural selection. Those few that are of known, but not hybrid, origin all seem to be the products of artificial stabilization processes brought about by human agency.
However, the reader should not suppose the foregoing facts imply hybridization typically produces agamosperms. The only claim made is that agamosperms are typically the products of hybridization. As we have repeatedly seen, hybridization can also produce non-agamospermous forms. Indeed, various crosses between plants capable of agamospermous reproduction give rise to sexual progeny. For example, Gustafsson (1946–1947: 145–146) says that the crosses Potentilla recta x P. adscharica and P. canescens x P. verna, which are both between agamosperms, yield F1 generations that are wholly or predominantly sexual. So hybridization can produce agamosperms from sexual parents and produce sexual progeny from agamospermous parents. The transition can proceed in either direction.
The important point is that agamosperms are a separate category of organism (in addition to polyploids) in which new forms of life are typically produced by stabilization processes associated with hybridization. NEXT PAGE >>